This is our second post about generating artificial data using Generative Adversarial Networks (GANs). For a broader introduction to the subject please refer to our previous article.
Our task at NeuroSYS was to build an image classifier for distinguishing between Petri dishes with bacteria or fungi grown in them and empty ones. In our dataset there were a lot of images containing multiple, large organisms, located close to the center of the dish and very few images with single, small organisms located near the edges. Both types of samples had to be assigned to the same category. i.e. samples containing organisms.
Below two types of samples are shown:
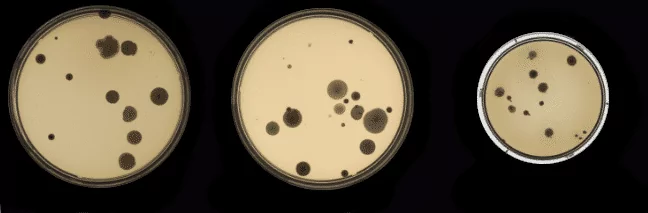

As a result, the classifier that we trained with this dataset recognized the first type of samples with high accuracy but the second type of samples was constantly classified as empty dishes. To solve this problem we decided to create artificial uncommon samples and use them to train a better classifier. We assumed that if classifier sees more untypical samples, it will learn to recognize them correctly.
First, we tried to apply a variation of standard GANs for the task. Unfortunately, we did not manage to generate real-looking images (for more detailed description please see our previous post).
In the second step, we decided to apply image-to-image translation with CycleGANs. Below we present the main idea behind this algorithm as well as the results that we were able to achieve with it.
Image-to-image translation
Image-to-image translation is a class of computer vision tasks where the goal is to learn how an input image can be mapped to an output image. For example, imagine that you took a picture of a view from your balcony and would like to see how it would be painted by Monet or Picasso. In this case the input image is your raw photo and the output image is the photo with painter’s style applied to it. This can be achieved by using a specific type of Generative Adversarial Networks (GANs), namely Cycle-Consistent Adversarial Network (CycleGAN) [1]. (Note: For a broader introduction to GANs please refer to our previous post).

How CycleGAN works
In order to train CycleGAN two sets of training data are needed, one set of images in domain X and the different set in domain Y. For example, the first set can contain photos of various landscapes and the other paintings of a certain artist. The goal is to learn to translate between the two domains, or in other words, to make photos look like paintings and vice versa. To do that CycleGAN uses two generators and two discriminators. One generatorGXY transforms images from domain X to Y and the other GYX converts images from Y domain to the X domain. Each generator has a corresponding discriminator DX and DY. As in standard GANs, discriminators try to correctly recognize fake and authentic data instances while generators aim at creating data instances, that will be deemed authentic by the discriminators.
In addition to that, two cycle consistency losses are introduced in order to push generators to be consistent with each other. They ensure, that if you translate an image from one domain to the other and back again by successively passing it through both generators you will get something similar to what you put in. The process is depicted in the figure below:
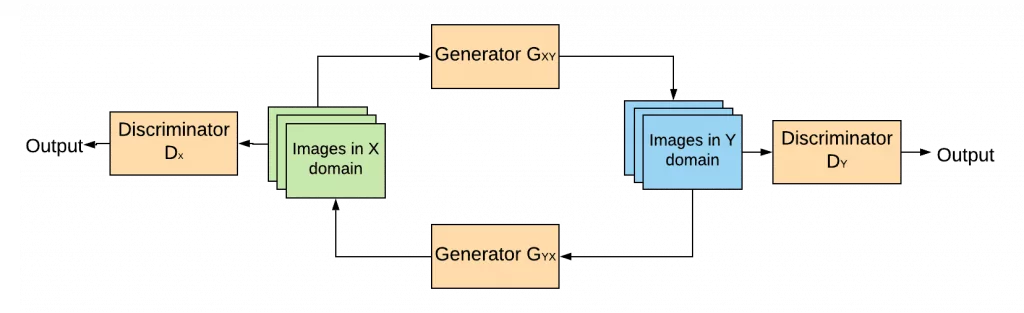
As for standard GANs, when CycleGAN is applied to visual data like images, the discriminator is a Convolutional Neural Network (CNN) that can categorize images and the generator is another CNN that learns a mapping from one image domain to the other.
Our results
We applied GANs to produce fake images of bacteria and fungi in Petri dishes. We aimed at creating samples that we lacked in the original training set, namely images with only few, small organisms located near the edges of the dish. First we used a variation of standard GANs called Improved Wasserstein GANs (for detailed description see our previous post).
Below samples generated by Improved Wasserstein GAN are presented:
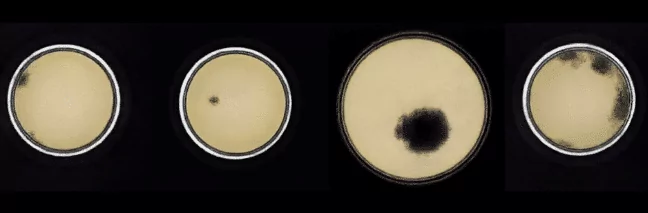
As you can see, we did not manage to create real-looking samples. We generated organisms of various sizes, both close to the center of the Petri dish and near the edges. However, bacteria and fungi on generated images do not look realistically and can be easily distinguished from the organisms present in the original set.
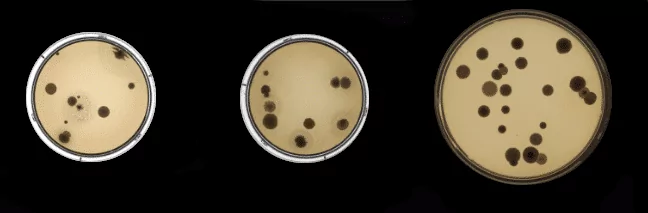
Then we decided to apply CycleGAN for the task. We constructed two sets. The first set was created by drawing black dots on images of empty Petri dishes. The other one contained real images of bacteria and fungi in Petri dishes. We hoped to transform dots into real looking organisms.
Below images in two domains are presented:

Below we present results that we obtained using CycleGAN. On the left side you can see empty dishes with dots from our training set and on the right side corresponding generated images:
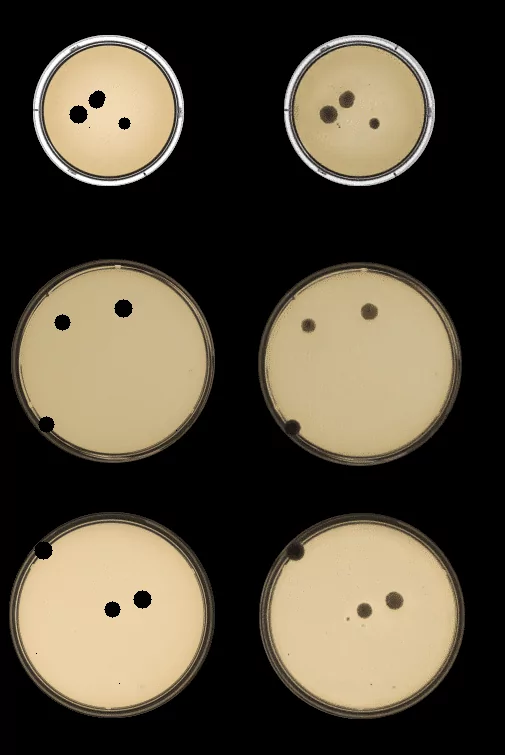
Using CycleGAN we managed to create real-looking samples. Generated bacteria and fungi are indistinguishable from the original ones. We can generate organisms of any size and at any location, because they arise exactly where the black dots were drawn and are of the same size. By drawing small dots near the edges of the Petri dish and transforming the images using trained CycleGAN we can create samples, that were scarce in our original dataset.
To sum up
We achieved our goal using CycleGANs. We were able to generate real-looking artificial samples that we lacked in the original training set and train classifier to correctly recognize untypical samples, namely images containing small organisms, located near the edges of the Petri dish.
References
[1] Jun-Yan Zhu, Taesung Park, Phillip Isola, and Alexei A Efros. Unpaired image-to-image translation using cycle-consistent adversarial networks. arXiv preprint arXiv:1703.10593, 2017
Project co-financed from European Union funds under the European Regional Development Funds as part of the Smart Growth Operational Programme.
Project implemented as part of the National Centre for Research and Development: Fast Track.